Current Projects
AI-based identification of cellular phenotypes in genetic screens
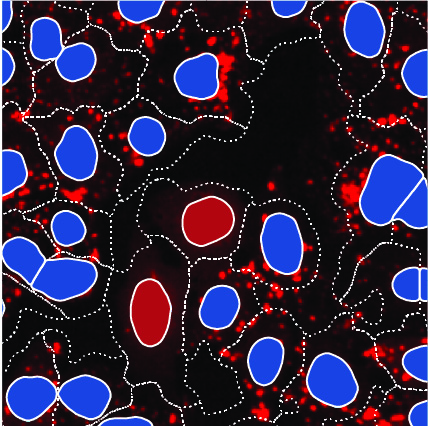

Using my SPARCS technology, I conduct image-based genetic screens in cellular model systems at human genome scale. These screens create datasets of hundreds of millions of single-cell fluorescence images. Identifying cells with relevant phenotypes in these datasets is a challenge.
I use self-supervised learning to train AI models to identify interesting phenotypes based on single-cell images. These models learn to identify specific genetic effects, enabling me to discover new gene functions.
Modeling cellular behavior across data modalities with scPortrait
Comprehensive computational modeling of a cell's responses to its surroundings — often called "virtual cell" — requires integrating different data modalities into a single model. One modality for which data can be acquired at a scale compatible with modern AI techniques is microscopy images.
Working with Sophia Mädler and the labs of Fabian Theis and Matthias Mann, I developed scPortrait, a software package that processes raw microscopy fields-of-view into standardized single-cell image datasets that can be integrated with other modalities acquired at the single cell level.
With scPortrait, we
- Mapped dissociated single-cell transcriptomics data to tissues using flow matching
- Identified a morphology-defined subset of immune cells that are associated with ovarian cancer
- Embedded single-cell images across modalities into transcriptome-based atlases
scPortrait Manuscript →
Modeling cell biology experiments with AI
Prophet is a general-purpose AI model for cellular phenotype prediction that generalizes by modeling each cell biology experiment as a composite of three elements:
- A cellular model system the experiment is conducted in
- A perturbation applied to that model system
- A readout that characterises the model systems's response
Image-based genetic screening with SPARCS
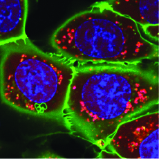
Together with Veit Hornung's and Matthias Mann's lab I developed the SPARCS technology to conduct image-based genetic screens in cell culture models with unprecedented throughput. In SPARCS, we image a library of genetically altered cells on a microscope, generate a single-cell image dataset using scPortrait and then select individual cells with interesting cells from hundreds of millions of single-cell images using AI-based cellular phenotyping methods. With its spatial resolution, SPARCS can screen complex phenotypes such as precise intracellular protein and organelle positions.
SPARCS enables a new type of genetic screens in which selective pressure is defined and applied by AI models. This leads to the discovery of entirely new biological mechanisms.
SPARCS Manuscript →